Global Zooplankton Biomass Distribution
R
Academic
Paper with Code
Analyzing and visualizing global zooplankton biomass patterns using R
This paper can be found here as well as the corresponding github repository.
Study Overview
- We used in situ imaging data from the Underwater Vision Profiler 5 (UVP5) to predict global zooplankton biomass distribution.
- The study covered a 10-year period (2008-2018) and analyzed 466,872 images from 3,549 stations worldwide.
- Boosted Regression Trees (BRTs) were used to model the relationship between zooplankton biomass and environmental variables.
Key Findings
- Global Biomass Estimate: The total zooplankton biomass in the upper 500m of the ocean was estimated at 0.403 Pg with an estimated 0.229 PgC for the epipelagic layer (0-200 m) and 0.173 PgC for the mesopelagic layer (200-500 m).
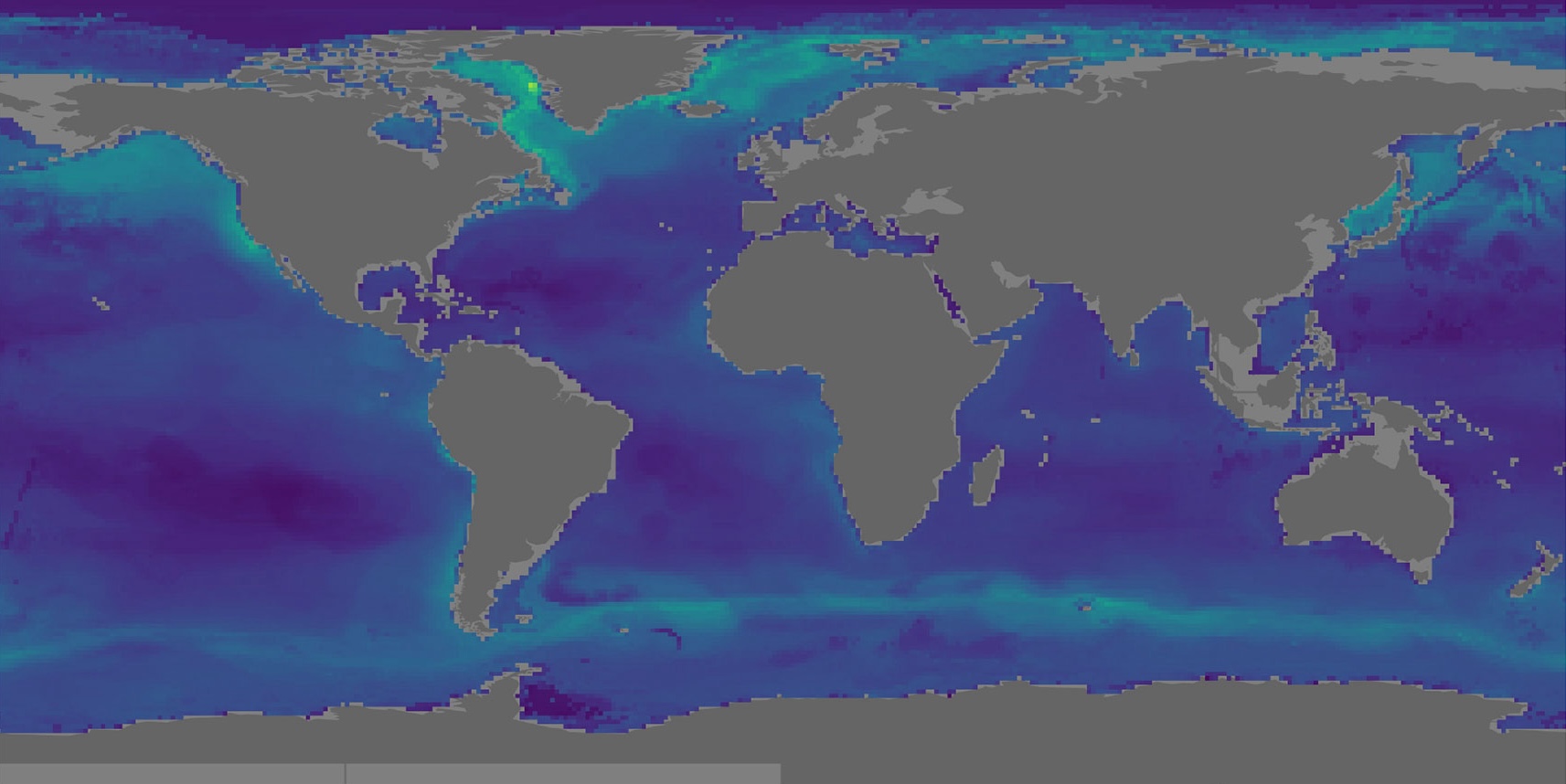
Distribution map of the predicted minimum global biomass between 0 and 500m (Figure 11 of the paper).
- Dominant Groups:
- Copepoda: 35.7% of total biomass
- Eumalacostraca: 26.6%
- Rhizaria: 16.4%
- Spatial Distribution:
- Highest biomass values were found around 60°N and 55°S.
- Lowest biomass was predicted in oceanic gyres and the Arctic (north of 80°N).
- An increase in biomass was observed around the equator.
Implications
- This study provides a new method for estimating global zooplankton biomass using in situ imaging and machine learning.
- The findings emphasize the importance of using non-intrusive sampling methods to accurately assess the biomass of fragile organisms like Rhizaria which has been underestimated in previous net-based studies.
- The results can be valuable for improving global biogeochemical models and understanding the role of zooplankton in marine ecosystems and the carbon cycle.
Perspectives
- Continued use of imaging systems like UVP5 and UVP6 to expand the dataset and improve predictions.
- Focus on sampling underrepresented areas, particularly in polar regions during winter.
- Integration of data from multiple imaging systems to cover a wider size spectrum of zooplankton.